Adeno-associated virus (AAV) vectors are important delivery platforms for therapeutic genome editing but are severely constrained by cargo limits. Simultaneous delivery of multiple vectors can limit dose and efficacy and increase safety risks. Here, we describe single-vector, ~4.8-kb AAV platforms that express Nme2Cas9 and either two sgRNAs for segmental deletions, or a single sgRNA with a homology-directed repair (HDR) template. We also use anti-CRISPR proteins to enable production of vectors that self-inactivate via Nme2Cas9 cleavage. We further introduce a nanopore-based sequencing platform that is designed to profile rAAV genomes and serves as a quality control measure for vector homogeneity. We demonstrate that these platforms can effectively treat two disease models [type I hereditary tyrosinemia (HT-I) and mucopolysaccharidosis type I (MPS-I)] in mice by HDR-based correction of the disease allele. These results will enable the engineering of single-vector AAVs that can achieve diverse therapeutic genome editing outcomes. Long-term expression of Cas9 following precision genome editing in vivo may lead to undesirable consequences. Here we show that a single-vector, self-inactivating AAV system containing Cas9 nuclease, guide, and DNA donor can use homology-directed repair to correct disease mutations in vivo.
Research and publish the best content.
Get Started for FREE
Sign up with Facebook Sign up with X
I don't have a Facebook or a X account
Already have an account: Login
 Your new post is loading... Your new post is loading...
 Your new post is loading... Your new post is loading...
|
|


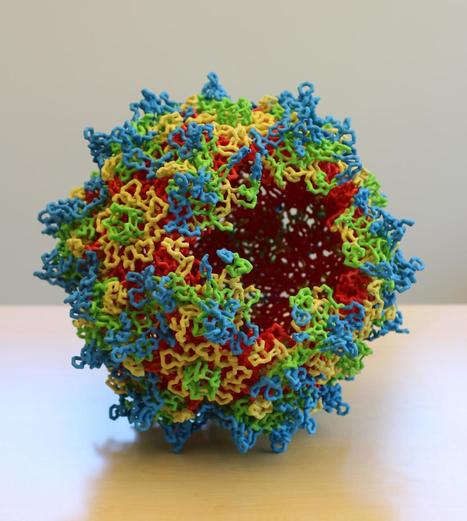





Most studies using AAVs to deliver gene therapy attempt to insert a Cas9 nuclease and a single guide RNA (sgRNA), along with their respective promoters, into the vector to create a single double-stranded break in the DNA. Given the 4.7 kb packaging limit of most AAVs, this is typically a significant challenge, especially for studies aimed at correcting genetic mutations by providing a donor DNA template for HDR. In this study, members of Professor Erik Sontheimer's lab at the University of Massachusetts Chan Medical School demonstrated the use of a cleverly designed, self-inactivating AAV vector with a compact Cas9 variant for therapeutic CRISPR genome editing in vivo. The study takes this packaging problem a step further, with the goal of creating a vector with enough cargo space to hold two sgRNAs rather than just one. In doing so, CRISPR could be used to excise pathogenic trinucleotide repeat expansions, such as that which causes Friedreich's ataxia. With the compact size of Staphylococcus aureus Cas9 (Nme1Cas9), this makes it easier to package into AAVs. It has a dinucleotide PAM sequence, which means it has a wider range of genomic targets and it displays high editing efficiencies in mammalian cells with low off-target activity, making it ideal for therapeutic applications.